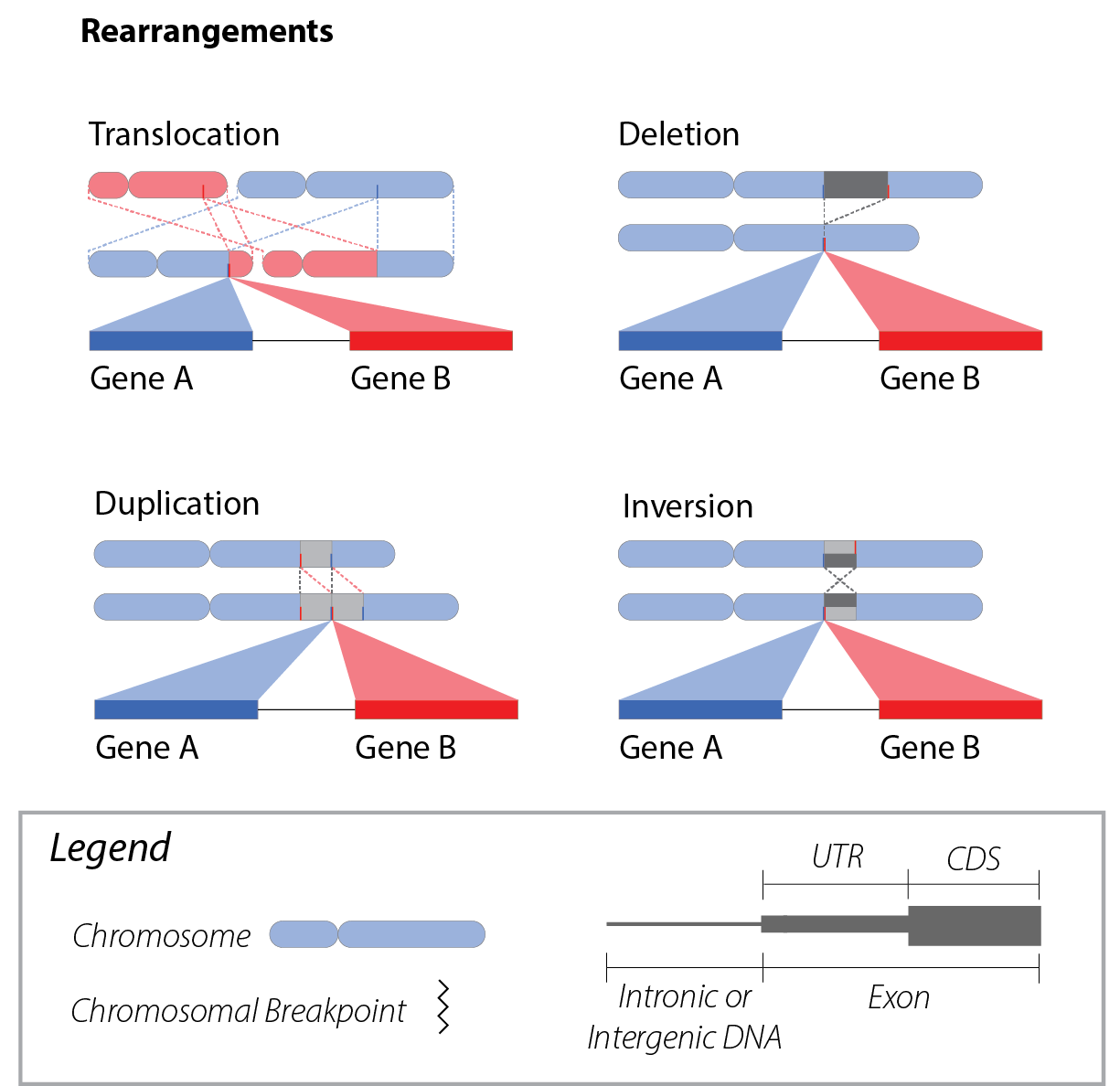
Terminology
Gene Fusions
Gene fusions are a complex class of variation that may be characterized by a broad range of relevant attributes with varying specificity. The defining characteristic of gene fusions is the interaction of two or more genes to drive aberrant activity of a gene product, through formation of a chimeric transcript or interaction of rearranged gene regulatory elements. Similar genetic variations involving Rearrangements within the same gene (e.g. internal tandem duplications), and transcript alterations due to splice site variants are biologically meaningful but distinct from gene fusions. Importantly, gene fusions are also distinct from the underlying genomic rearrangements that drive them, though these concepts have been conflated due to the historical use of genomic assays for inferring the presence of specific gene fusions.
The two primary classes of gene fusions–Chimeric Transcript Fusions and Regulatory Fusions–are not mutually exclusive classes, as some fusions (such as promoter-swap fusions) may be defined either in the context of their regulatory elements or by their chimeric gene product.
Gene products that are considered loss-of-function or are not expressed should not be described as gene fusions, even when they result from a genomic rearrangement.
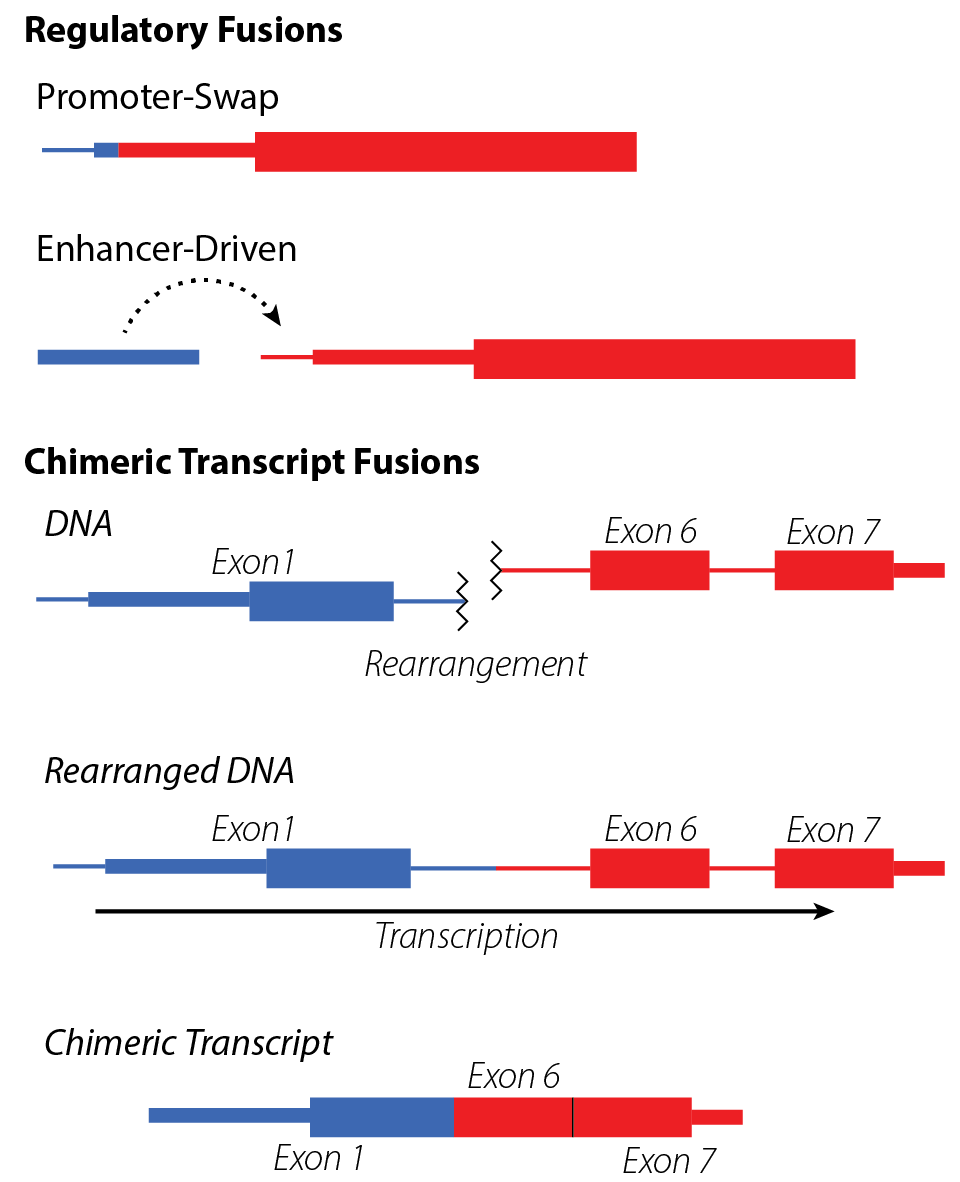
(TOP) Gene fusions may be regulatory in nature, where a rearranged promoter or nearby enhancer element drives overexpression of the partner gene. (BOTTOM) Gene fusions typically result in chimeric transcripts between two genes, which (for coding transcripts) often result in novel protein sequences.
Chimeric Transcript Fusions
Chimeric transcript gene fusions are often driven by genomic rearrangements involving two gene loci, resulting in the concatenation of exons from each into a single chimeric transcript. This class of fusions is exemplified by well-known clinically-relevant gene fusions such as BCR(hgnc:1014)::ABL1(hgnc:76). Other clinically-relevant gene fusions of this type may be driven by RNA processing events in lieu of genomic rearrangements, including read-through derived fusions such as CTSD(hgnc:2529)::IFITM10(hgnc:40022) and trans-splicing derived fusions such as JAZF1(hgnc:28917)::JJAZ1(hgnc:17101). These alternative mechanisms for creating fusions are specified in these guidelines, but it should be noted that most read-through and trans-splicing events are artifactual and have little to no known clinical relevance.
Regulatory Fusions
In contrast to chimeric transcript fusions, deregulated gene fusions are primarily characterized by the rearrangement of regulatory elements from one gene near a second gene, resulting in the increased gene product expression of the second gene. This class of gene fusions include promoter-swapping gene fusions such as TMPRSS2(hgnc:11876)::ERG(hgnc:3446), as well as enhancer-driven gene fusions such as Reg@IGH(hgnc:5477)::MYC(hgnc:7553). Gene products rendered unexpressed or non-functional should not be described as gene fusions, even when they result from a genomic rearrangement.
Gene Fusion Contexts
Determining the salient elements for a gene fusion is dependent upon the context in which the gene fusion is being described, whether it describes an assayed fusion event from a sample (Assayed Gene Fusions) or an aggregate context described in biomedical literature or knowledgebases (Categorical Gene Fusions). These guidelines provide recommendations for characterizing gene fusions in each context.
Assayed Gene Fusions
Assayed gene fusions from biological specimens are directly detected using RNA-based gene fusion assays, or alternatively may be inferred from genomic rearrangements detected by whole genome sequencing or by coarser-scale cytogenomic assays in the context of informative phenotypic biomarkers. For example, an EWSR1 fusion is often inferred by breakapart FISH assay when a neoplasm is diagnosed or suspected to be Ewing sarcoma/primitive neuroectodermal tumor by immunohistochemical and/or morphological analysis.
Categorical Gene Fusions
In contrast, categorical gene fusions are generalized concepts representing a class of fusions by their shared attributes, such as retained or lost regulatory elements and/or functional domains, and are typically curated from the biomedical literature for use in genomic knowledgebases. An example categorical gene fusion is EWSR1 as a known 5’ gene fusion partner that binds 3’ partner genes with DNA-binding domains.